There was lots of discussion on twitter yesterday about Graham Coop’s paper with Peter Ralph (or vice versa), on The geography of recent genetic ancestry across Europe, particularly regarding the FAQ they’d created.
I was eager to take a look, and, it’s slightly embarrassing to say, I first did a search to see if they’d made a connection to any of my work. (I’m probably not the only one to do that.) Sure enough, they cited a paper of mine, but it was Giglo et al. (2001) Am J Hum Genet 68: 874–883, on “my” chr 8p inversion, and not what I’d expected, my autozygosity paper.
What did the chr 8p inversion have to do with this? Search for “[36]” and you’ll find:
We find that the local density of IBD blocks of all lengths is relatively constant across the genome, but in certain regions the length distribution is systematically perturbed (see Figure S1), including around certain centromeres and the large inversion on chromosome 8 [36], also seen by [35].
The chr 8p inversion presents an interesting data analysis story from my postdoc years. In a nutshell: I was studying human crossover interference, found poor model fit for maternal chr 8 that was due to tight apparent triple-crossovers in two individuals in each of two families, hypothesized that there was an inversion in the region, but it would have to be both long and with both orientations being common. The inversion was confirmed via FISH, and it’s something like 5 Mbp long, with the frequencies of the two orientations being 40 and 60% in people of European ancestry.
Marshfield maps and crossover interference
I was a postdoc with Jim Weber in Marshfield, Wisconsin, 1997-1999. My main effort concerned the construction of human genetic maps using data from eight of the CEPH families. I was particularly interested in characterizing crossover interference.
I found that the gamma model fit the data quite well. These are histograms of the inter-crossover distances, with expected distributions for different models:
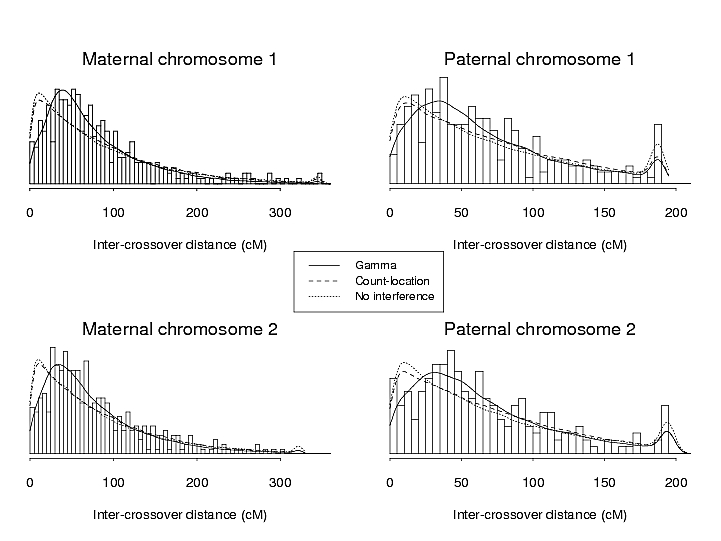
But the model fit poorly for maternal chromosome 8:
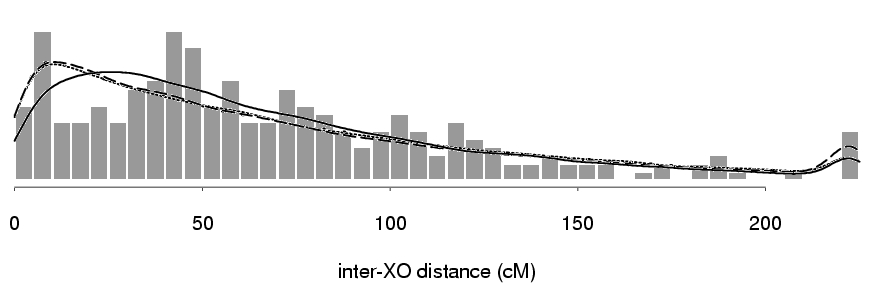
Why the poor model fit?
Why was the model fit for maternal chromosome 8 so terrible? It turned out that there was a set of four tight apparent triple-crossovers, two in each of two families. The black and white dots indicate grandmother and grandfather DNA on different meiotic products:
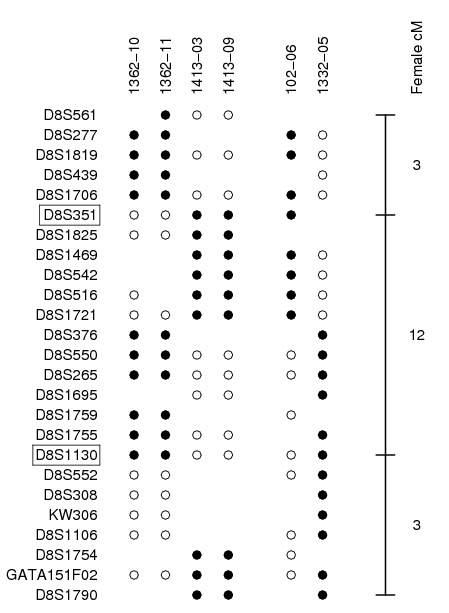
I saw these tight triple-crossovers and thought, “Oops! I got the marker order wrong.” (Remember, this was before we had a physical map.) But if you reverse the orientation of the region, crossovers in other individuals would become triple-crossovers.
So I thought, perhaps this is an inversion polymorphism: some individuals have the region in one orientation and others have it in the opposite orientation. But it would have to be long (it was an estimated 12 cM in females: something like 5 Mbp). And both orientations would have to be common, since they would each need to be present in the homozygous state for there to be recombination events.
The inversion
Jim Weber contacted David Ledbetter, and folks in his group investigated the region and confirmed, via FISH, that there was indeed a long, common inversion polymorphisms on chromosome 8p.
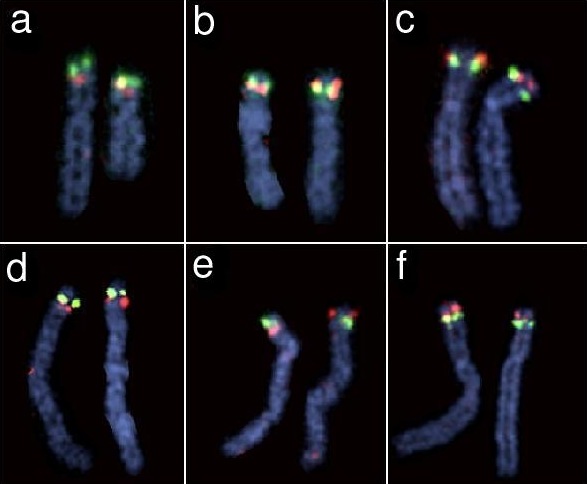
They marked one side of the region in green and the other side red, and in the left panels green is above red on both chromosomes, in the right panels red is above green on both chromosomes, and in the center there is one chromosome with each orientation. Analysis of further subjects indicated that the two orientations have allele frequencies 40 and 60% in people of European ancestry.
Authorship order
I referred to this story in passing in the past. A fortuitous clinical connection was made to this chr 8p inversion, and due a dispute over author order, what should have been one paper got split into two, and my half trickled down the journal chain to finally appear in Terry Speed’s Festschrift.
I try not to discuss author order anymore. I care only about presence/absence.
A related story: autozygosity
This wasn’t the only surprising finding to come from my efforts on the Marshfield genetic maps.
An important part of the map construction was data cleaning: identifying tight double-crossovers indicative of genotyping errors. I looked at piles of CRI-MAP chrompic output, to find such double crossovers. Here’s a somewhat nicer image:
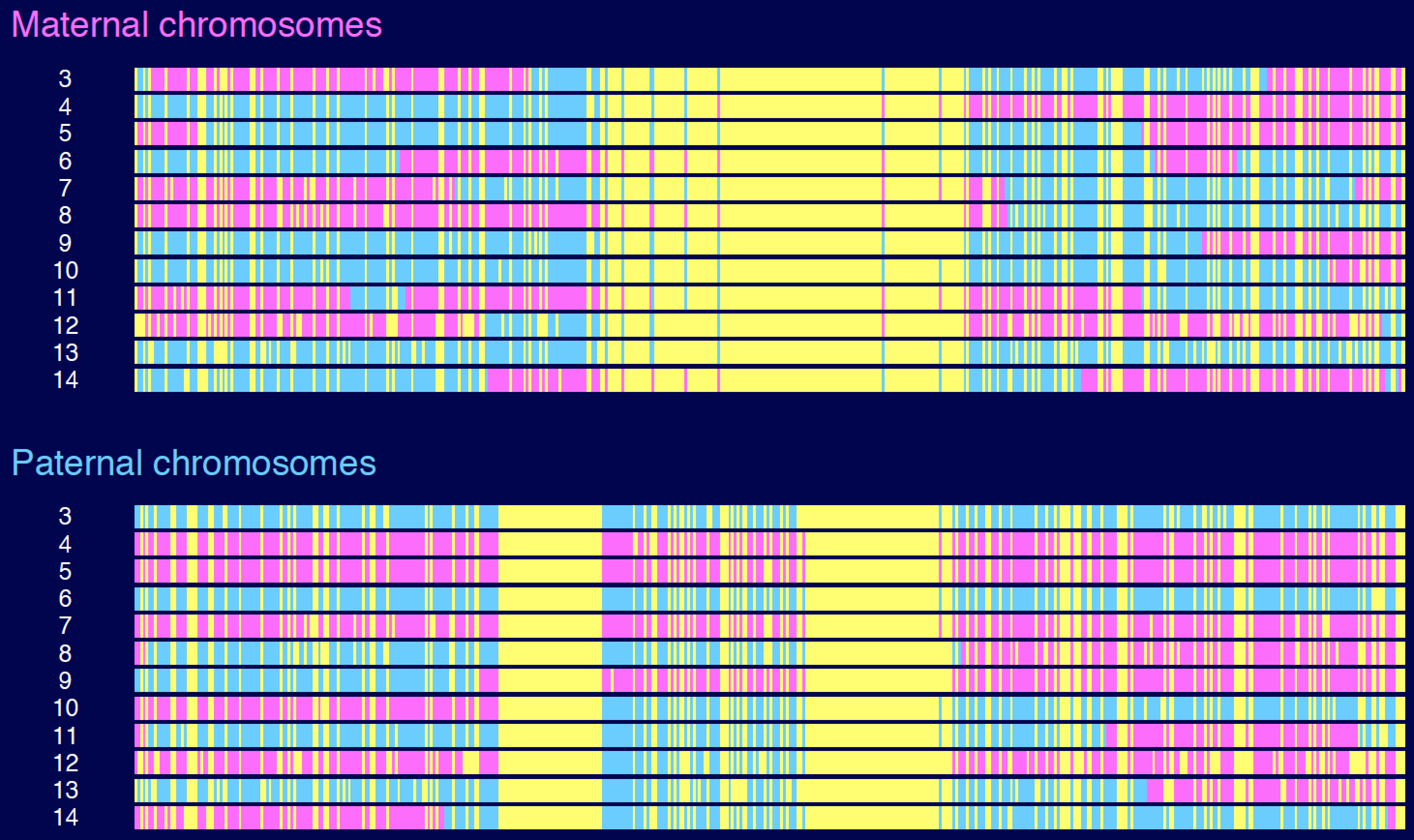

Pink and blue indicate grandmother and grandfather DNA, respectively, on the maternal and paternal chromosomes in each individual from a large sibship. Yellow indicates missing data: if the mother or father was homozygous, the grandparental origin of DNA was indeterminate. Why these long stretches of homozygosity?
After publishing the genetic map paper, I went back to study these long stretches of homozygosity. And then I looked for such regions more systematically, and found lots of them:
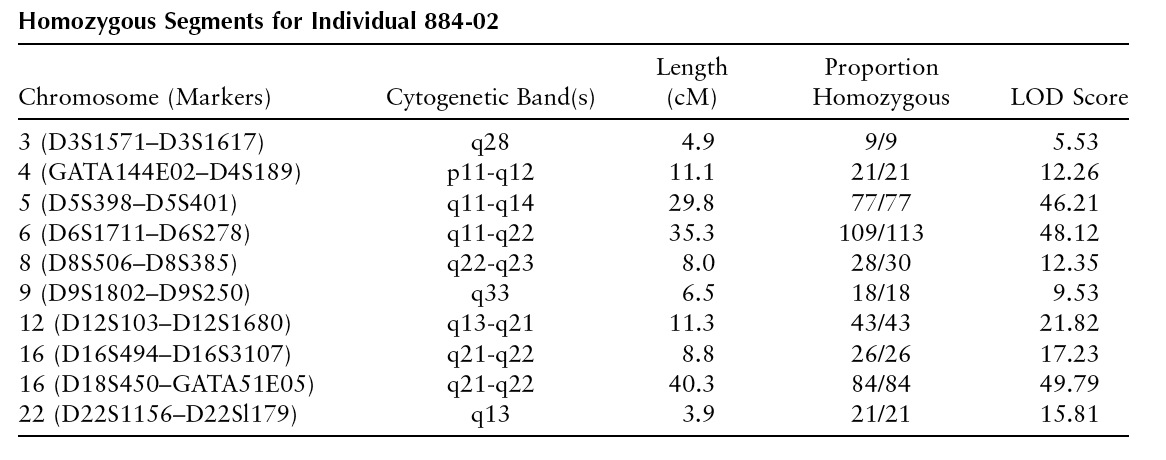
It turned out to be autozygosity: in two CEPH families, the grandparents were related, and so the parents where homozygous by descent (aka autozygous) for chunks of their genome.
Moral
In both of these cases, apparent artifacts in the data led to the most interesting findings in the work. If your model doesn’t fit, or you see something odd, ask, “Why is that?”