One of the coolest R packages I heard about at the useR! Conference: Toby Dylan Hocking’s directlabels package for putting labels directly next to the relevant curves or point clouds in a figure.
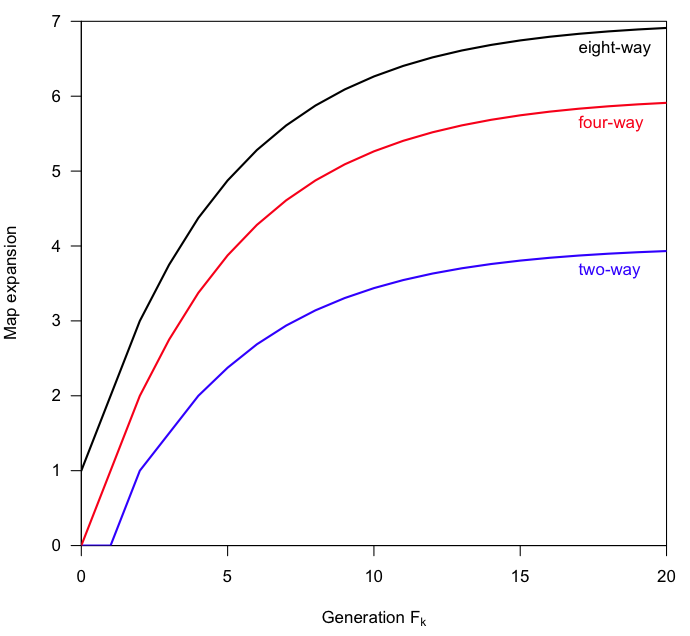
I think I first learned about this idea from Andrew Gelman: that a separate legend requires a lot of back-and-forth glances, so it’s better to put the labels right by the relevant bits. For example, like this:
rather than this:
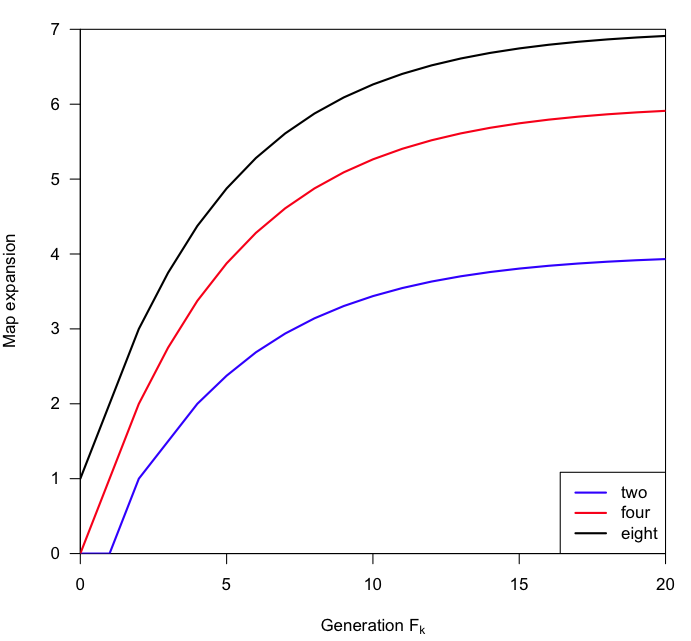
I’ve adopted this approach as much as possible, though it often requires a bit of work (and thought) to get the labels in just the right place. Here’s the code I used for the first of those pictures. (It was relatively easy here.)
load(con <- url("https://www.biostat.wisc.edu/~kbroman/blog/mapexpansion.RData"))
close(con)
plot(dat[,1], dat[,4],
xlab=expression(paste("Generation ", F[k])),
ylab="Map expansion",
type="l", lwd=2, col="black", las=1,
xaxs="i", yaxs="i", ylim=c(0, 7))
for(i in 2:3)
lines(dat[,1], dat[,i], lwd=2, col=c("blue", "red")[i-1])
text(17, dat[18,-1]-0.1, paste(colnames(dat)[-1], "-way", sep=""),
adj=c(0,1), col=c("blue","red","black"))
The aim of the directlabels package is get this without effort.
I need to switch to either lattice or ggplot2 (vs. base graphics). But I should be doing that anyway. I’ll try lattice for this. I rearrange the data a bit, call xyplot and then use direct.labels to make the actual plot, as follows. (Note the use of with and gl, which I just learned about from Richie Cotton.)
library(lattice)
library(directlabels)
dat2 <- with(dat, data.frame(gen=rep(gen,3),
mapexpansion=c(two, four, eight),
cross=gl(3, nrow(dat), labels=c("two","four","eight"))))
p <- xyplot(mapexpansion ~ gen, data=dat2, groups=cross, type="l", lwd=2)
direct.label(p)
And here’s what I get:
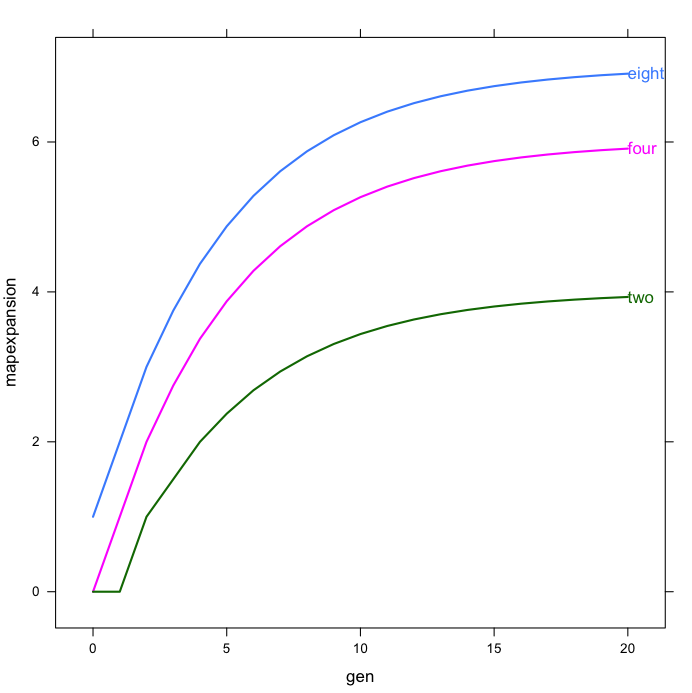
For a final figure for publication, one will want to do some editing by hand, but for day-to-day graphics, this looks really useful. The following is the “real” version of the above figure, from a paper under review, using a mixture of legend and direct labels:
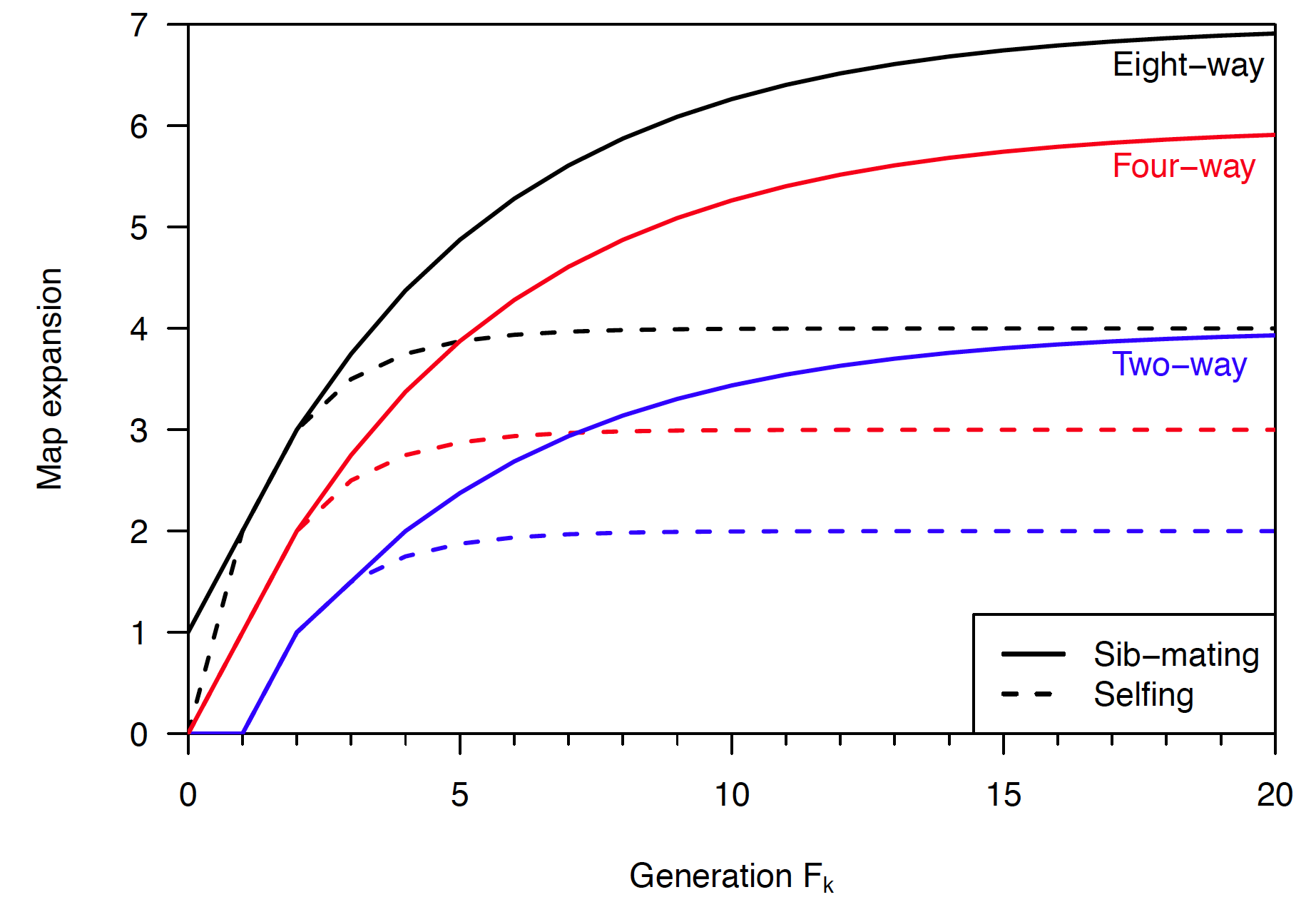
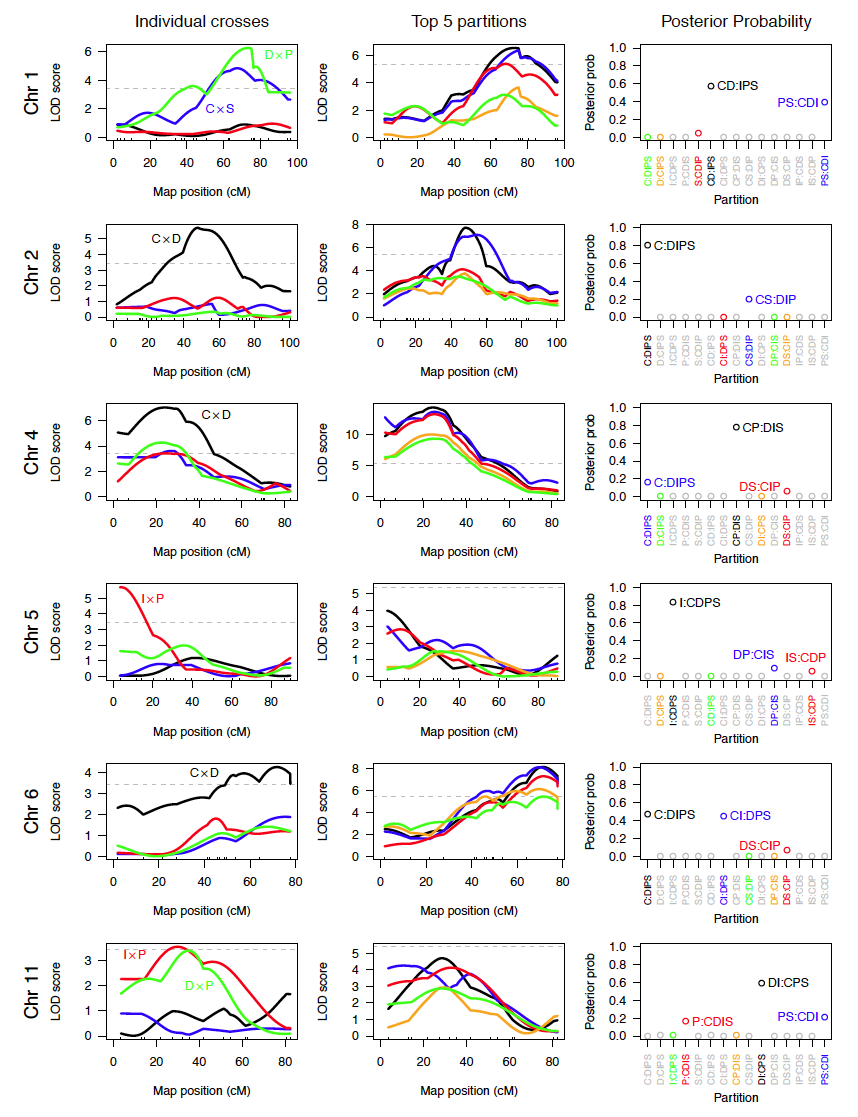
Here’s another figure I’m quite proud of, from a paper nearing submission.